- 1Université Paris-Saclay, CNRS, GEOPS, 91405, Orsay, France
- 2Belgian Institute for Space Aeronomy (BIRA-IASB), Avenue Circulaire, 3 B-1180 Brussels Belgium
- 3School of Physical Sciences, The Open University, Milton Keynes, MK7 6AA, U.K.
- 4INAF-Istituto di Astrofisica e Planetologia Spaziali, Rome, ITALY
- 5Instituto de Astrofísica de Andalucia CSIC
Abstract
One of the main difficulties to analyze modern spectroscopic datasets is the extremely large amount of data. For example, in atmospheric transmittance spectroscopy, the solar occultation channel (SO) of the NOMAD instrument onboard the ESA ExoMars2016 satellite called Trace Gas Orbiter (TGO) had produced ~10 millions of spectra in ~20000 acquisition sequences since the beginning of the mission in April 2018 until 15 January 2020. Usually, new lines are discovered after a long iterative process of model fitting and manual residual analysis.
Here we propose a new method, based on unsupervised machine learning, to automatically detect new minor species. Although precise quantification is out of scope, this tool can also be used to quickly summarize the dataset.
The methodology is the following: first we suggest to approximate the dataset by a linear mixture of abundance and endmember spectra. Then, unsupervised source separation is used, in form of non-negative matrix factorization. Several methods are tested on synthetic and simulation data.
On synthetic example, this approach is able to detect chemical compounds present at 1.5 times the noise level for 100 hidden spectra out of 104. Results on simulated spectra of NOMAD-SO targeting CH4 show a detection limit of 100 ppt in favorable conditions. Results on real martian data from NOMAD-SO show that CO2 and H2O are present, as expected, but CH4 is absent. Nevertheless, we find a set of new unexpected lines in the database.
Dataset
We propose here to focus on the Nadir and Occultation for MArs Discovery (NOMAD) instrument and especially the Solar Occultation (SO) channel [1], operating at wavenumbers from 2320 cm−1 to 4550 cm−1 (wavelength 2.2 to 4.3 μm).
Method
We propose to simplify the non-linear radiative transfer into a linear mixture. The collection of observation X is approximated by a few sources S and each of them present with an abundances A.
X=A.S (1)
Several algorithms have been proposed to solve this problem, subject to positivity (both S and A are non-negative). Such problem is called Non negative Matrix Factorization (NMF) [2]. This constraint is important to keep the physical meaning, but also to promote sparsity of S (a signal is sparse when a lot of values are close to zero except several non-zero values).
Results
Figure 1 and 2 present a synthetic toy example and demonstrate the capability of the method to extract a pure CH4 contribution, even hidden – 100 out of 10000 at 3-σ level of the noise.
Figure 3 illustrates the results on NOMAD data for order 136. Separated contribution of H2O and background are identified. No endmembers seemed to be related to CH4.
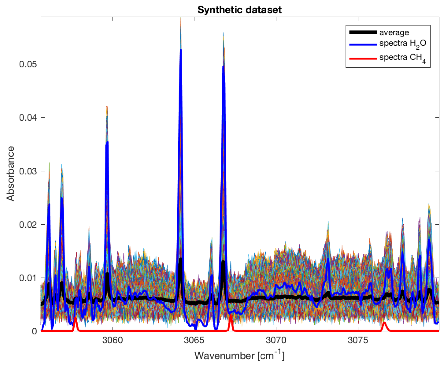
Figure 1: Synthetic dataset containing 104 spectra with various abundances of H2O and 100 containing CH4 at 3-σ level of the noise. In blue the reference spectra of H2O. In red the reference spectra of CH4.
Figure 2: Analysis of the dataset presented in figure 1 for NS = 4. Endmembers 1 and 3 are identified to the level with significant noise contribution, endmember 2 is identified to H2O, and endmember 4 is CH4.
Figure 3: (top) Results for real data of order 136 for NS = 5. The endmember 1 is identified to the level background (continuum misestimation), the endmembers 2, 3 and 4 are identified to H2O, either directly either from the adjacent orders. No endmember seems to be related to CH4. (bottom) Synthetic spectra from PSG [3]
Conclusion
We proposed a new machine learning tool [4], based on non-negative matrix factorization, to automatically detect new minor species. We applied it on potential CH4 detection on NOMAD-SO but future work should also focus on other order to allow new discovery by serendipity. Our tool may also be applied on other planetary spectral dataset, such as surface measurement.
Acknowledgements
We acknowledge support from the “Institut National des Sciences de l’Univers” (INSU), the "Centre National de la Recherche Scientifique" (CNRS) and "Centre National d’Etudes Spatiales" (CNES) through the "Programme National de Planétologie" and the ExoMars TGO programs. The NOMAD experiment is led by the Royal Belgian Institute for Space Aeronomy (BIRA-IASB), assisted by Co-PI teams from Spain (IAA-CSIC), Italy (INAF-IAPS), and the United Kingdom (Open University). This project acknowledges funding by the Belgian Science Policy Office (BELSPO), with the financial and contractual co-ordination by the ESA Prodex Office (PEA 4000103401, 4000121493), by Spanish Ministry of Science and Innovation (MCIU) and by European funds under grants PGC2018-101836-B-I00 and ESP2017-87143-R (MINECO/FEDER), as well as by UK Space Agency through grants ST/R005761/1, ST/P001262/1, ST/R001405/1 and ST/R001405/1 and Italian Space Agency through grant 2018-2-HH.0. This work was supported by the Belgian Fonds de la Recherche Scientifique - FNRS under grant number 30442502 (ET-HOME). The IAA/CSIC team acknowledges financial support from the State Agency for Research of the Spanish MCIU through the Center of Excellence Severo Ochoa award for the Instituto de Astrofísica de Andalucía (SEV-2017-0709).
References
[1] Vandaele et al., Space Science Reviews 214, 5, 2018
[2] Lee, D. D., Seung, H. S., 401, 788-791, Nature, 1999.
[3] Villanueva et al., 217, 86-104, JQSRT, 2018
[4] Schmidt et al., under review in JQSRT, 2020
How to cite: Schmidt, F., Cruz Mermy, G., Erwin, J., Robert, S., Neary, L., Thomas, I., Daerden, F., Ristic, B., Patel, M., Bellucci, G., Lopez-Moreno, J.-J., and Vandaele, A. C.: Machine learning for automatic identification of new minor species, Europlanet Science Congress 2020, online, 21 Sep–9 Oct 2020, EPSC2020-781, https://doi.org/10.5194/epsc2020-781, 2020.

